Nextflow
Description
Nextflow is free open source software distributed under the Apache 2.0 licence developed by Seqera Labs. The project was started in the Notredame Lab at the Centre for Genomic Regulation (CRG).
The source code for Nextflow is available on GitHub.
Architecture
There are four components of a Nextflow engine as deployed in an Amazon Genomics CLI context:
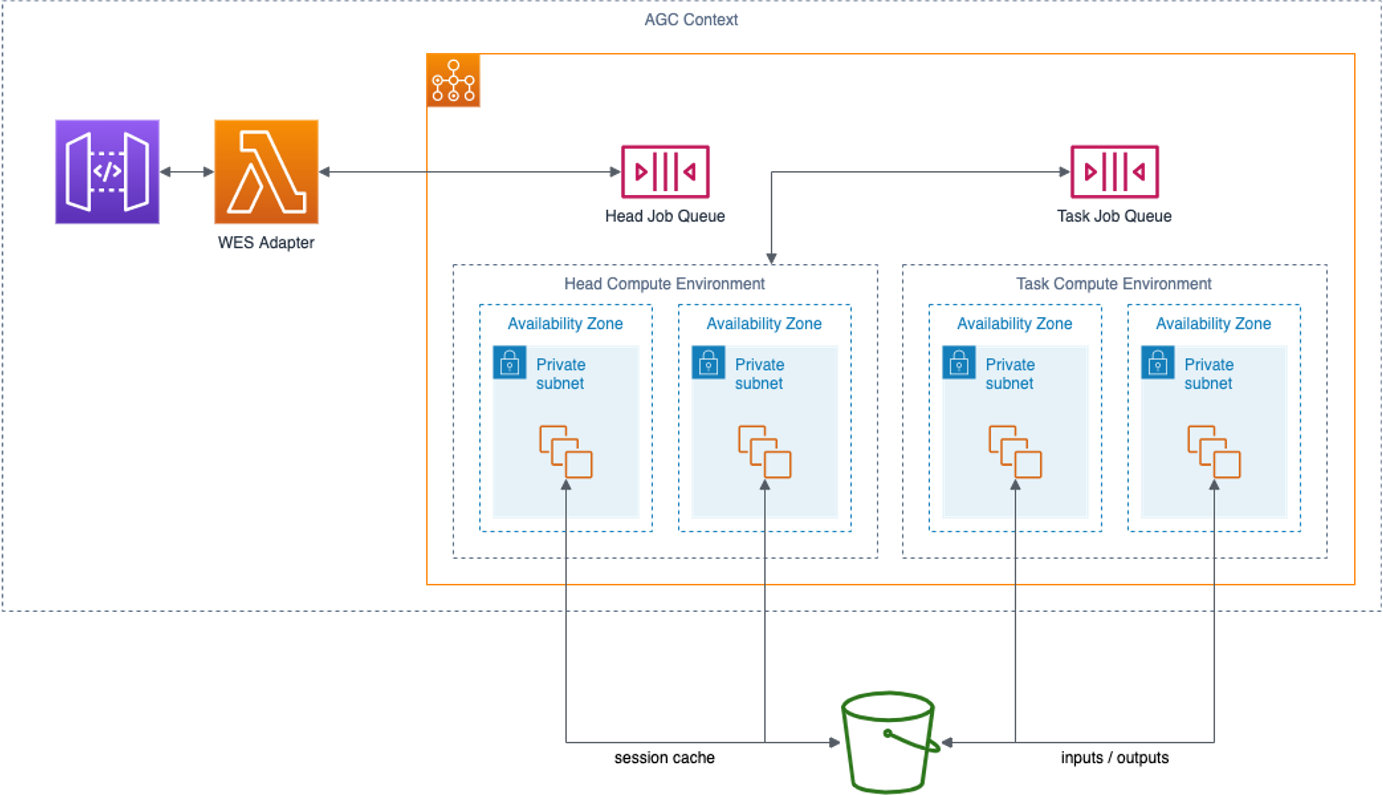
WES Adapter
Amazon Genomics CLI communicates with the Nextflow engine via a GA4GH WES REST service. The WES Adapter implements the WES standard and translates WES calls into calls to the Nextflow head process.
Head Compute Environment
For every workflow submitted, the WES adapter will create a new AWS Batch Job that contains the Nextflow process responsible for running that workflow. These Nextflow “head” jobs are run in an “On-demand” compute environment even when the actual workflow tasks run in a Spot environment. This is to prevent Spot interruptions from terminating the workflow coordinator.
Task Compute Environment
Workflow tasks are submitted by the Nextflow head job to an AWS Batch queue and run in containers using an AWS Compute Environment. Container characteristics are defined by the resources requested in the workflow configuration. AWS Batch coordinates the elastic provisioning of EC2 instances (container hosts) based on the available work in the queue. Batch will place containers on container hosts as space allows.
Fetch and Run Strategy
Execution of workflow tasks uses a “Fetch and Run” strategy. Input files required by a workflow task are fetched from S3 into the task container. Output files are copied out of the container to S3.
Disk Expansion
Container hosts in the Batch compute environment use EBS volumes as local scratch space. As an EBS volume approaches a capacity threshold, new EBS volumes will be attached and merged into the file system. These volumes are destroyed when AWS Batch terminates the container host.